Abstract
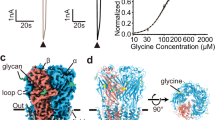
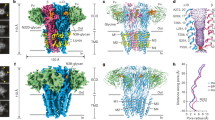
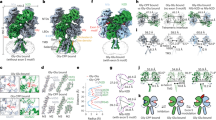
Cys-loop receptors are pentameric ligand-gated ion channels (pLGICs) that mediate fast synaptic transmission. Here functional pentameric assembly of truncated fragments comprising the ligand-binding N-terminal ectodomains and the first three transmembrane helices, M1–M3, of both the inhibitory glycine receptor (GlyR) α1 and the 5HT3A receptor subunits was found to be rescued by coexpressing the complementary fourth transmembrane helix, M4. Alanine scanning identified multiple aromatic residues in M1, M3 and M4 as key determinants of GlyR assembly. Homology modeling and molecular dynamics simulations revealed that these residues define an interhelical aromatic network, which we propose determines the geometry of M1–M4 tetrahelical packing such that nascent pLGIC subunits must adopt a closed fivefold symmetry. Because pLGIC ectodomains form random nonstoichiometric oligomers, proper pentameric assembly apparently depends on intersubunit interactions between extracellular domains and intrasubunit interactions between transmembrane segments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Liao, M.J., Huang, K.S. & Khorana, H.G. Regeneration of native bacteriorhodopsin structure from fragments. J. Biol. Chem. 259, 4200–4204 (1984).
Popot, J.L., Gerchman, S.E. & Engelman, D.M. Refolding of bacteriorhodopsin in lipid bilayers. A thermodynamically controlled two-stage process. J. Mol. Biol. 198, 655–676 (1987).
Mackenzie, K.R. Folding and stability of α-helical integral membrane proteins. Chem. Rev. 106, 1931–1977 (2006).
Popot, J.L. & Engelman, D.M. Membrane protein folding and oligomerization: the two-stage model. Biochemistry 29, 4031–4037 (1990).
Popot, J.L. & Engelman, D.M. Helical membrane protein folding, stability, and evolution. Annu. Rev. Biochem. 69, 881–922 (2000).
de Planque, M.R. & Killian, J.A. Protein-lipid interactions studied with designed transmembrane peptides: role of hydrophobic matching and interfacial anchoring. Mol. Membr. Biol. 20, 271–284 (2003).
Eilers, M., Shekar, S.C., Shieh, T., Smith, S.O. & Fleming, P.J. Internal packing of helical membrane proteins. Proc. Natl. Acad. Sci. USA 97, 5796–5801 (2000).
Liu, Y., Gerstein, M. & Engelman, D.M. Transmembrane protein domains rarely use covalent domain recombination as an evolutionary mechanism. Proc. Natl. Acad. Sci. USA 101, 3495–3497 (2004).
Bowie, J.U. Solving the membrane protein folding problem. Nature 438, 581–589 (2005).
Hildebrand, P.W. et al. Hydrogen-bonding and packing features of membrane proteins: functional implications. Biophys. J. 94, 1945–1953 (2008).
Gimpelev, M., Forrest, L.R., Murray, D. & Honig, B. Helical packing patterns in membrane and soluble proteins. Biophys. J. 87, 4075–4086 (2004).
Hildebrand, P.W., Rother, K., Goede, A., Preissner, R. & Frommel, C. Molecular packing and packing defects in helical membrane proteins. Biophys. J. 88, 1970–1977 (2005).
Hall, Z.W. Recognition domains in assembly of oligomeric membrane proteins. Trends Cell Biol. 2, 66–68 (1992).
Green, W.N. & Millar, N.S. Ion-channel assembly. Trends Neurosci. 18, 280–287 (1995).
Gu, Y., Camacho, P., Gardner, P. & Hall, Z.W. Identification of two amino acid residues in the epsilon subunit that promote mammalian muscle acetylcholine receptor assembly in COS cells. Neuron 6, 879–887 (1991).
Chavez, R.A., Maloof, J., Beeson, D., Newsom-Davis, J. & Hall, Z.W. Subunit folding and αδ heterodimer formation in the assembly of the nicotinic acetylcholine receptor. Comparison of the mouse and human α subunits. J. Biol. Chem. 267, 23028–23034 (1992).
Kuhse, J., Laube, B., Magalei, D. & Betz, H. Assembly of the inhibitory glycine receptor: identification of amino acid sequence motifs governing subunit stoichiometry. Neuron 11, 1049–1056 (1993).
Griffon, N. et al. Molecular determinants of glycine receptor subunit assembly. EMBO J. 18, 4711–4721 (1999).
Nicke, A. et al. P2X1 and P2X3 receptors form stable trimers: a novel structural motif of ligand-gated ion channels. EMBO J. 17, 3016–3028 (1998).
Nicke, A., Rettinger, J., Mutschler, E. & Schmalzing, G. Blue native PAGE as a useful method for the analysis of the assembly of distinct combinations of nicotinic acetylcholine receptor subunits. J. Recept. Signal Transduct. Res. 19, 493–507 (1999).
Büttner, C. et al. Ubiquitination precedes internalization and proteolytic cleavage of plasma membrane-bound glycine receptors. J. Biol. Chem. 276, 42978–42985 (2001).
Sadtler, S. et al. A basic cluster determines topology of the cytoplasmic M3–M4 loop of the glycine receptor α1 subunit. J. Biol. Chem. 278, 16782–16790 (2003).
Nicke, A., Thurau, H., Sadtler, S., Rettinger, J. & Schmalzing, G. Assembly of nicotinic α7 subunits in Xenopus oocytes is partially blocked at the tetramer level. FEBS Lett. 575, 52–58 (2004).
Gendreau, S. et al. A trimeric quaternary structure is conserved in bacterial and human glutamate transporters. J. Biol. Chem. 279, 39505–39512 (2004).
Detro-Dassen, S. et al. Conserved dimeric subunit stoichiometry of SLC26 multifunctional anion exchangers. J. Biol. Chem. 283, 4177–4188 (2008).
Duckwitz, W., Hausmann, R., Aschrafi, A. & Schmalzing, G. P2X5 subunit assembly requires scaffolding by the second transmembrane domain and a conserved aspartate. J. Biol. Chem. 281, 39561–39572 (2006).
Green, T., Stauffer, K.A. & Lummis, S.C. Expression of recombinant homo-oligomeric 5-hydroxytryptamine3 receptors provides new insights into their maturation and structure. J. Biol. Chem. 270, 6056–6061 (1995).
Claudio, T. et al. Fibroblasts transfected with Torpedo acetylcholine receptor β-, γ-, and δ-subunit cDNAs express functional receptors when infected with a retroviral α recombinant. J. Cell Biol. 108, 2277–2290 (1989).
Blount, P. & Merlie, J.P. Mutational analysis of muscle nicotinic acetylcholine receptor subunit assembly. J. Cell Biol. 111, 2613–2622 (1990).
Buck, T.M., Wagner, J., Grund, S. & Skach, W.R. A novel tripartite motif involved in aquaporin topogenesis, monomer folding and tetramerization. Nat. Struct. Mol. Biol. 14, 762–769 (2007).
Schmidt, T.G., Koepke, J., Frank, R. & Skerra, A. Molecular interaction between the Strep-tag affinity peptide and its cognate target, streptavidin. J. Mol. Biol. 255, 753–766 (1996).
Hilf, R.J. & Dutzler, R. X-ray structure of a prokaryotic pentameric ligand-gated ion channel. Nature 452, 375–379 (2008).
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4Å resolution. J. Mol. Biol. 346, 967–989 (2005).
Hilf, R.J. & Dutzler, R. Structure of a potentially open state of a proton-activated pentameric ligand-gated ion channel. Nature 457, 115–118 (2009).
Bocquet, N. et al. X-ray structure of a pentameric ligand-gated ion channel in an apparently open conformation. Nature 457, 111–114 (2009).
McGaughey, G.B., Gagne, M. & Rappe, A.K. π-Stacking interactions. Alive and well in proteins. J. Biol. Chem. 273, 15458–15463 (1998).
Gurezka, R., Laage, R., Brosig, B. & Langosch, D. A heptad motif of leucine residues found in membrane proteins can drive self-assembly of artificial transmembrane segments. J. Biol. Chem. 274, 9265–9270 (1999).
Lemmon, M.A., Treutlein, H.R., Adams, P.D., Brunger, A.T. & Engelman, D.M. A dimerization motif for transmembrane α-helices. Nat. Struct. Biol. 1, 157–163 (1994).
Rath, A. & Deber, C.M. Surface recognition elements of membrane protein oligomerization. Proteins 70, 786–793 (2008).
Choma, C., Gratkowski, H., Lear, J.D. & Degrado, W.F. Asparagine-mediated self-association of a model transmembrane helix. Nat. Struct. Biol. 7, 161–166 (2000).
Zhou, F.X., Cocco, M.J., Russ, W.P., Brunger, A.T. & Engelman, D.M. Interhelical hydrogen bonding drives strong interactions in membrane proteins. Nat. Struct. Biol. 7, 154–160 (2000).
Burley, S.K. & Petsko, G.A. Aromatic-aromatic interaction: a mechanism of protein structure stabilization. Science 229, 23–28 (1985).
Gervasio, F.L., Chelli, R., Procacci, P. & Schettino, V. The nature of intermolecular interactions between aromatic amino acid residues. Proteins 48, 117–125 (2002).
Ridder, A., Skupjen, P., Unterreitmeier, S. & Langosch, D. Tryptophan supports interaction of transmembrane helices. J. Mol. Biol. 354, 894–902 (2005).
Johnson, R.M., Hecht, K. & Deber, C.M. Aromatic and cation-π interactions enhance helix-helix association in a membrane environment. Biochemistry 46, 9208–9214 (2007).
Sal-Man, N., Gerber, D., Bloch, I. & Shai, Y. Specificity in transmembrane helix-helix interactions mediated by aromatic residues. J. Biol. Chem. 282, 19753–19761 (2007).
Li, S.C. & Deber, C.M. A measure of helical propensity for amino acids in membrane environments. Nat. Struct. Biol. 1, 368–373 (1994).
Nooren, I.M. & Thornton, J.M. Diversity of protein-protein interactions. EMBO J. 22, 3486–3492 (2003).
Grudzinska, J. et al. The β subunit determines the ligand binding properties of synaptic glycine receptors. Neuron 45, 727–739 (2005).
Gloor, S., Pongs, O. & Schmalzing, G. A vector for the synthesis of cRNAs encoding Myc epitope-tagged proteins in Xenopus laevis oocytes. Gene 160, 213–217 (1995).
Becker, D. et al. The P2X7 carboxyl tail is a regulatory module of P2X7 receptor channel activity. J. Biol. Chem. 283, 25725–25734 (2008).
Schägger, H., Cramer, W.A. & von Jagow, G. Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal. Biochem. 217, 220–230 (1994).
Schägger, H. & von Jagow, G. Tricine-sodium dodecycl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 166, 368–379 (1987).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Eswar, N., Eramian, D., Webb, B., Shen, M.Y. & Sali, A. Protein structure modeling with MODELLER. Methods Mol. Biol. 426, 145–159 (2008).
Brejc, K. et al. Crystal structure of an ACh-binding protein reveals the ligand-binding domain of nicotinic receptors. Nature 411, 269–276 (2001).
Brooks, B.R. et al. CHARMM: A program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983).
Pettersen, E.F. et al. UCSF Chimera—visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Morris, G.M. & Lim-Wilby, M. Molecular docking. Methods Mol. Biol. 443, 365–382 (2008).
Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008).
Acknowledgements
We thank U. Braam for expert help in generating cDNA constructs, G. Drwal for electrophysiological recording and F. Pult for sharing with us the data of her M.D. work on C-terminal N-glycosylation. Supported by Deutsche Forschungsgemeinschaft (La1086/5-3; Schm536/4-3), Fonds der Chemischen Industrie (H.B.), the Hertie Foundation (B.L.), the Alexander von Humboldt Foundation (H.B., V.T.), Deutscher Akademischer Austauschdienst (D.K.) and the Russian Academy of Sciences grant “Molecular and Cell Biology” (V.T., D.K.).
Author information
Authors and Affiliations
Contributions
S.H. designed and generated DNA constructs and performed biochemical experiments; D.K. performed homology modeling and molecular dynamics simulations and analyzed the data; S.D.-D. synthesized cRNAs, generated DNA constructs and performed biochemical experiments; N.L. generated mutants and performed biochemical experiments; M.K. performed electrophysiological recordings; V.T. supervised homology modeling and analyzed data; H.B. designed experiments and wrote the paper; B.L. designed experiments, performed electrophysiological recordings, analyzed data and wrote the paper; G.S. designed constructs and biochemical experiments, analyzed the data and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 2821 kb)
Rights and permissions
About this article
Cite this article
Haeger, S., Kuzmin, D., Detro-Dassen, S. et al. An intramembrane aromatic network determines pentameric assembly of Cys-loop receptors. Nat Struct Mol Biol 17, 90–98 (2010). https://doi.org/10.1038/nsmb.1721
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1721
This article is cited by
-
BAC transgenic mice to study the expression of P2X2 and P2Y1 receptors
Purinergic Signalling (2021)
-
Thermophoretic analysis of ligand-specific conformational states of the inhibitory glycine receptor embedded in copolymer nanodiscs
Scientific Reports (2020)
-
A lipid site shapes the agonist response of a pentameric ligand-gated ion channel
Nature Chemical Biology (2019)
-
The roles of aromatic residues in the glycine receptor transmembrane domain
BMC Neuroscience (2018)
-
An allosteric link connecting the lipid-protein interface to the gating of the nicotinic acetylcholine receptor
Scientific Reports (2018)