Key Points
-
The zebrafish has become a widely used model organism because of its fecundity, its morphological and physiological similarity to mammals, the existence of many genomic tools and the ease with which large phenotype-based screens can be performed. These attributes might also enable efficiency increases in several steps in the drug development process.
-
Large-scale genetic and morpholino oligonucleotide screens allow unbiased discovery of genes that cause a desired phenotype. Zebrafish screens for genetic or epigenetic perturbations that suppress a disease phenotype can be used to discover novel therapeutic targets.
-
Historically, numerous drugs have been discovered by observing phenotypic changes in whole animals exposed to small molecules, but these discoveries have often been serendipitous. Large-scale, systematic screens can be performed in the zebrafish to identify small molecules that can suppress disease phenotypes.
-
At present, assessment of toxicity often occurs independently from efforts to discover lead compounds and improve their potency. High-throughput zebrafish toxicity assays combine many of the advantages of in vitro and in vivo toxicity models, making it possible to assess toxicity much earlier in the drug development process.
Abstract
The zebrafish has become a widely used model organism because of its fecundity, its morphological and physiological similarity to mammals, the existence of many genomic tools and the ease with which large, phenotype-based screens can be performed. Because of these attributes, the zebrafish might also provide opportunities to accelerate the process of drug discovery. By combining the scale and throughput of in vitro screens with the physiological complexity of animal studies, the zebrafish promises to contribute to several aspects of the drug development process, including target identification, disease modelling, lead discovery and toxicology.
Similar content being viewed by others
Main
Drug discovery involves a complex iterative process of biochemical and cellular assays, with final validation in animal models, and ultimately in humans. Mammalian models of absorption, distribution, metabolism and excretion (ADME)/pharmacokinetics and efficacy are expensive, laborious and consume large quantities of precious compounds. There is also increasing pressure to limit animal use to situations in which they are absolutely necessary, such as in preclinical toxicity and safety assessment. Zebrafish are beginning to be used at various stages of the drug discovery process and can be a useful and cost-effective alternative to some mammalian models (such as rodents, dogs and pigs). In this article, we will review the use of zebrafish in target validation, disease modelling, target and lead compound discovery, and toxicology.
Zebrafish background
The zebrafish has been a popular pet for decades. Its use for research increased substantially approximately 8 years ago, following the demonstration that it is amenable to large-scale forward genetic screens1. Such large-scale genetic screens had previously been limited to invertebrates such as flies, worms and yeast. Zebrafish made it possible to take a forward genetic approach to understanding vertebrate-specific processes that affect development and disease. In the past decade, thousands of mutations have been generated that affect ORGANOGENESIS, physiology and behaviour2,3. These mutations have proved to be a rich source of information about the relationships between genes and function.
In the past few years, several additional tools have been developed that have greatly increased the utility of the zebrafish as an experimental organism (Table 1). The zebrafish genome has now been sequenced by the Sanger Center, and there has been substantial annotation of the genome through the trans-National Institutes of Health Zebrafish Genome Initiative. Collections of full-length zebrafish cDNAs are available, as are multiple DNA microarrays for expression-profiling experiments4,5. Gene expression can also be rapidly analysed using whole-embryo in situ hybridization. Gene function can be rapidly and robustly studied in zebrafish using antisense MORPHOLINO OLIGONUCLEOTIDES6,7. Furthermore, techniques for generating transgenic lines8,9,10, targeted mutations (reverse genetics)11 and cloning by nuclear transfer12 have been developed. Together, these tools have made the zebrafish the organism of choice for many researchers, as demonstrated by the marked increase in zebrafish publications in recent years. The yearly number of PubMed references on zebrafish has almost tripled in the past 5 years and increased more than tenfold in the past decade (Fig. 1a).
a | The increase in the use of zebrafish for research is depicted as the number of zebrafish-related Pubmed references for each year from 1980 to 2003. The term 'zebrafish' was used for searching Pubmed. b–e | The transparent body of the zebrafish larva is shown 5 days post-fertilization. Images of increasing magnification show the ability to discern structures on the systemic (c), organ (d, heart shown) and cellular (e, tail shown) scales in the living organism. f | Specific structures can be highlighted in the living zebrafish by transgenic expression of fluorescent proteins from tissue-specific promoters. Here, endothelial cells are marked by expression of enhanced green fluorescent protein from the fli1 (friend leukemia integration 1) promoter. g | Two zebrafish embryos developing in a single well of a standard 384-well assay plate.
Beyond genetics and experimental tools, the strength of the zebrafish resides in the analysis of phenotype. Perhaps no other organism (and certainly no vertebrate) is better suited to high-throughput phenotyping. The zebrafish embryo is optically transparent, making it possible to detect functional and morphological changes in internal organs without having to kill or dissect the organism (Fig. 1b–e). These functional and morphological changes can be further highlighted by the use of transgenic lines and other reporter molecules (Fig. 1f). For example, fluorescent labelling of MAUTHNER CELLS was recently used to identify conditions that promote post-injury regeneration of damaged zebrafish neurons13. In another example, fluorescently quenched phospholipids were used to detect lipid processing and transport in live, intact zebrafish14. The transparent fish and fluorescent marker were combined to facilitate the rapid phenotyping of thousands of individuals in a large-scale screen for genes that affect lipid trafficking.
The scale that can be achieved in zebrafish experiments is impressive by vertebrate standards. Early zebrafish embryos are less than 1 mm in diameter, allowing several embryos to fit easily in a single well of a 384-well plate (Fig. 1g). What they lack in size they make up for in numbers — each female can lay up to 300 eggs at a time. Even a small zebrafish facility can generate many thousands of embryos at a time. Miniature size, large numbers and easy detection of a phenotype of interest are the sine qua non of high-throughput screening (HTS) and, indeed, zebrafish have proved to be well suited to HTS, as described in Fig. 2.
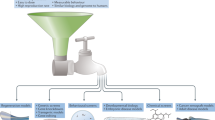
a | Adult zebrafish are mated, producing 200–300 embryos per female. b | Embryos are distributed to 96- or 384-well assay plates. Typically, three embryos are placed in each well. c | A small-molecule library that contains potentially biologically active compounds is synthesized or acquired. d | Small molecules are added to the water surrounding the zebrafish. A single small molecule or combinations can be added to each well. e | After a period of incubation, the phenotypic effects of the small molecules on the zebrafish are determined visually or through an automated read-out. f | Referral to the library database reveals the identities of the biologically active small molecules.
Identifying novel drug targets
Identifying and validating novel drug targets remains a bottleneck in drug discovery15. Despite a wealth of information about normal physiology and disease pathology, it is still difficult to predict which targets will effectively reverse a disease phenotype without producing unwanted side effects. Consequently, drug discovery efforts directed at poorly validated targets frequently fall short of expectations16,17.
Genetic and morpholino oligonucleotide screens in the zebrafish are an efficient means of systematically assessing the roles of individual genes in disease processes. As such, they represent a promising route to the identification and validation of novel drug targets. In traditional forward genetic zebrafish screens, phenotypes have been identified that resemble human diseases18,19,20. The novel genes that underlie the zebrafish disease phenotypes might lead directly to the identification of novel drug targets. Alternatively, the disease phenotypes might form the basis of further screens to find genes or chemicals that can correct the phenotype.
Genetic screens
During the past decade, numerous forward genetic screens have been carried out in zebrafish. Thousands of distinct mutations have been identified, and more than 400 of them have been cloned2,3,21. So far, the emphasis of most of these screens has been on developmental biology. Therefore, although these screens have provided a tremendous amount of information about the roles of genes during embryogenesis, the relevance of most of the identified genes to disease pathology remains poorly established. However, several disease-related screens have been performed, including the following examples.
Polycystic kidney disease. Forward genetic screens have been used to identify several mutations that cause kidney cysts in zebrafish analogous to those found in humans with polycystic kidney disease (PKD)22,23. The mutated genes include pkd2 and vhnf1 (also known as transcription factor 2; tcf2 ), genes that are implicated in human PKD, as well as several novel genes. Many of the identified genes are involved in cilia function, lending strong support to the emerging hypothesis the PKD is caused by ciliary defects.
Cholesterol processing. A genetic screen has been used to identify zebrafish mutants with normal digestive organ morphology that have altered phospholipid and cholesterol processing14.
Tissue regeneration. Mutants have been identified that affect the ability to regenerate damaged tissues, including fins and hearts24.
Heart disease. Mutants identified through the use of zebrafish screens recapitulate several aspects of human heart disease. In several cases, the zebrafish and human disease states are caused by orthologous mutations. In both zebrafish and humans, TTN (titin) mutations cause cardiomyopathy25,26; TBX5 (T-box 5) mutations cause congenital heart defects27,28; and HERG (also known as potassium voltage-gated channel, subfamily H (ether a go go-related), member 2; KCNH2 ) mutations cause arrhythmias29,30.
Anaemias. The iron transporter ferroportin was first discovered in zebrafish as a mutant with embryonic anaemia31. Subsequently, it was found that ferroportin is defective in patients with HAEMOCHROMATOSIS TYPE 4 (Ref. 32). This is the first time that a zebrafish gene has led to the discovery of a new human disease gene. Also, the discovery of the kgg mutant, which lacks haematopoietic stem cells and is defective in the cdx–hox pathway, led to a new method for amplifying blood stem cells from mammalian embryonic stem cells33.
Cancer. It is possible to model cancer in the zebrafish using various transgenic fish. So far, there are excellent models of leukaemia and melanoma in the zebrafish20,34,35.
Nervous system. Zebrafish models of numerous nervous system disorders have been described, including retinal degeneration and anxiety36,37. In addition, genetic screens to identify mutations that cause deafness, blindness, mechanosensation defects, loss of CONDITIONED PLACE PREFERENCE and other behavioural defects have been reported38,39,40,41.
The forward genetic screens and models described above provide important entrance points into pathways that might contribute to numerous disease processes. Ultimately, genetic suppressor screens are likely to add great value. For example, starting with a disease model, it should be possible to screen for mutations that suppress or delay the development of the disease phenotype, with the mutated genes becoming logical drug target candidates. A similar approach in mice was recently reported, in which genetic suppressors of genetic THROMBOCYTOPAENIA were identified. Two mutant alleles of c-Myb were identified that suppress the disease phenotype, indicating that c-Myb might be a therapeutic target for thrombocytopaenia42. Genetic suppressor screens in zebrafish have not been reported, but all of the required elements for such screens are now available.
Morpholino oligonucleotide screens
Morpholinos are chemically modified antisense oligonucleotides that can be designed to hybridize to the translation-initiation or splicing acceptor/donor sites of specific mRNAs6. When injected into zebrafish embryos, they cause a robust knockdown of gene function. The developmental roles of many individual genes have been determined by injection of morpholinos in zebrafish.
Could morpholino oligonucleotide-mediated gene knockdown be used to discover novel therapeutic targets? One promising approach involves inducing a disease state in zebrafish (through the use of infectious agents, genetic mutations or pharmacological agents) combined with large-scale morpholino screening. By systematically knocking down many genes, it should be possible to identify gene knockdowns that prevent or slow the development of the disease phenotype. Small-scale morpholino screens have been reported in Xenopus tropicalis43 and Ciona intestinalis44, and zebrafish libraries containing morpholinos that target several hundred genes have been assembled. As the size and availability of these libraries increase, they are likely to be powerful tools for the systematic, unbiased identification of therapeutic targets.
Therapeutic targets identified by forward genetic or morpholino oligonucleotide screens could become the focus of conventional, target-based drug discovery efforts, including in vitro HTS. Alternatively, small-molecule screens to identify suppressors of zebrafish disease phenotypes can be initiated before the identification of a validated target, as described below.
Phenotype-based target and lead discovery
When a therapeutic target has been identified and validated, in vitro HTS based on target binding or function can often be used to identify novel structures that modify the activity of the target protein. Alternatively, medicinal chemistry can frequently be used to generate novel derivatives of compounds that are known to affect the target. For example, since the discovery of the 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase inhibitor mevastatin, dozens of structurally related statins have been developed45.
What about diseases for which the optimal therapeutic targets have not yet been identified? At present, validated targets represent only a small fraction of potential targets, and developing therapies for many of the most significant diseases (including atherosclerosis, chronic myelogenous leukaemia and bronchoalveolar lung cancer) is limited by the fact that effective targets have not yet been identified for these diseases. Predicting which proteins must be targeted to improve a disease state remains surprisingly difficult and, consequently, selecting novel targets for drug development on the basis of in vitro data entails significant risk16,17. In many cases, drug development directed at novel targets has proceeded smoothly at early in vitro stages but has failed at late stages, either because disruption of the target did not produce the expected effect in whole animals or because it also caused unexpected side effects. Clearly, many in vitro enzymatic assays are poor surrogates for complex physiological diseases. It is perhaps not surprising, therefore, that developing novel therapies guided solely by in vitro assays often leads to unexpected results in whole organisms.
One alternative to in vitro target-based drug discovery is discovery that is guided by phenotype in the context of a whole organism. Whereas target-based approaches can lead to the discovery of compounds that modify a target but that might not modify the disease, phenotype-based approaches can be used to discover compounds that modify the disease phenotype, regardless of the specific molecular target. The development of many drugs in use today was guided by phenotype in whole organisms (Table 2). For example, to discover the novel cholesterol-lowering drug ezetimibe (Zetia), scientists at Schering-Plough initially used a target-based approach46. They predicted that inhibitors of acyl-coenzyme A cholesterol acyltransferase (ACAT) would lead to a reduction in serum cholesterol, and so they developed in vitro assays for screening for ACAT inhibitors. Although potent, specific inhibitors of ACAT were identified, none of the compounds was effective at reducing serum cholesterol in the cholesterol-fed hamster. Remarkably, in the process of screening through the ineffective ACAT inhibitors, a synthetic byproduct was identified that reduced serum cholesterol in the hamsters. Guided solely by phenotype — namely serum cholesterol levels in the hamster — the byproduct was identified46. Derivatives of the byproduct were synthesized and tested for efficacy — again guided by phenotype alone — resulting finally in the development and Food and Drug Administration (FDA) approval of ezetimibe in 2002. The precise target of ezetimibe remains unknown, although it seems to block cholesterol absorption in the intestine47.
Because the mechanism of action of ezetimibe is distinct from other classes of cholesterol-lowering drug, such as the statins, it is a powerful additional tool for treating hypercholesterolaemia. In this case, the target-based approach to the development of ACAT inhibitors was unsuccessful, whereas the phenotype-based approach enabled the discovery of a novel cholesterol-lowering drug that functions through a novel and unexpected mechanism of action.
The examples described above illustrate the utility of being able to follow the phenotypic effects of compounds in a physiologically relevant, organismal context. However, systematizing this approach can be difficult. In the case of ezetimibe, a serendipitous observation sparked further experimentation, which necessitated the use of many animals and painstaking phenotyping46. Without the benefits of serendipity, a large-scale phenotypic screen could theoretically have been performed but would have required even more animals and more phenotyping. In one such Herculean screen, Squibb scientists performed a whole-animal, phenotype-based screen for antituberculosis agents. They screened 5,000 compounds in mice, en route to the successful phenotype-based discovery of isoniazid48.
Enter the zebrafish. Despite the demonstrated potential of this approach to discover novel, effective therapies, the resources, space and effort required to carry out this type of screen have prohibited their widespread use. To be practical, phenotype-based whole-organism screens need to be much less expensive, consume smaller quantities of compounds, take up less space and use simpler phenotyping than the mammalian assays described above. All of these requirements can be met by screens in whole zebrafish. The feasibility of performing phenotype-based small-molecule screens in zebrafish is illustrated by two recent examples in which chemical suppressors of specific genetic mutations were identified. Zebrafish that have the gridlock mutation, which causes a vascular defect, and crb (crash and burn), a cell-cycle mutation, were used to screen through diverse small-molecule libraries. In both screens, small molecules that can reverse the phenotypic effects of the mutations were identified49,50.
A screen for suppressors of the gridlock mutation. Gridlock mutants have a hypomorphic mutation in the hey2 gene, a hairy/enhancer of split-related transcriptional repressor believed to function downstream of Notch in regulating vascular cell-fate decisions51,52. Mutant embryos develop a dysmorphogenesis of the dorsal aorta that prevents circulation to the trunk and tail, although perfusion of the head is normal. Mutant embryos were distributed into 96-well plates, with three embryos per well, and compounds from a structurally diverse small-molecule library were pin-transferred into the water surrounding the zebrafish embryos, with one compound per well. The embryos were allowed to develop and visually scored for the gridlock phenotype under a dissecting microscope. After screening 5,000 small molecules, two structurally related compounds were identified that completely suppress the gridlock mutant phenotype, causing mutant embryos that are exposed to the compounds to develop a normal vasculature (Fig. 3)49. The gridlock-suppressing compounds, called GS4012 and GS3999, represent a novel class of compounds that were not previously known to influence vasculogenesis or angiogenesis. The compounds seem to rescue the aortic defect directly rather than promoting the formation of collateral vessels. No ectopic vessels have been observed in treated embryos (R.T.P., unpublished observations), which go on to survive through adulthood. Their mechanism of action is not yet known, although they seem to activate the vascular endothelial growth factor (VEGF) pathway, and overexpression of VEGF is sufficient to rescue the gridlock mutation49.
The perfusion of tissues with blood has a significant role in many diseases, including myocardial infarction, stroke, diabetes and cancer. Identifying novel classes of compounds and novel mechanisms for regulating the formation of blood vessels might have therapeutic applications. As initial evidence that the gridlock suppressors might have vasculogenic activity in a mammalian system, GS4012 was found to promote the formation of endothelial tubules from human cells in an in vitro matrigel assay49.
A screen for a suppressor of the crb mutant phenotype. An embryo-based small-molecule screen was performed to identify suppressors of the crb mutant phenotype50. crb mutants have an increased number of mitotic cells, and the heterozygotes have an increased rate of cancer. A total of 100 pairs of crb heterozygotes generated enough embryos to screen 1,000 compounds per week at 20 μM each. The entire DIVERSet E library (Chembridge, San Diego), which contains 16,320 compounds, was screened in 16 weeks. Because embryos were generated by mating heterozygous crb mutants, the embryos used for the screen had a phenotypic ratio of three wild-type embryos to one mutant embryo. Three classes of compound that had different effects on wild-type and mutant embryos were identified. Class I included 21 small molecules that increased phosphorylated histone H3 (pH3) staining in both wild-type and mutant embryos, and class II consisted of 11 chemicals that decreased pH3 staining in all embryos (R. Murphey, H. M. Stern and L.I.Z., manuscript in preparation). A third class consisted of one compound that specifically suppressed the elevated pH3 staining of crb embryos and had little effect on wild-type embryos. Because of its ability to suppress a specific cell-cycle defect in mutant cells without affecting wild-type cells, this compound might be useful as an anticancer agent.
Structure–activity relationships
The zebrafish as a system is highly amenable to the study of structure–activity relationships (SARs). During the above screen for compounds that alter pH3, several compounds were identified that perturb the cell cycle, but do so in both wild-type and mutant embryos. SARs have been established for several of these compounds. As shown in Fig. 4, several derivatives of parent compound L were generated and tested for their ability to alter pH3 levels in intact zebrafish. Three compounds induce the zebrafish phenotype at concentrations similar to those of the parent compound, four compounds induce the phenotype only at a fivefold higher concentration, and three compounds have no activity (R. Murphey and L.I.Z., unpublished observations).
Compound L was identified in a zebrafish assay for changes in phosphohistone H3 levels. Testing derivatives of L in zebrafish rapidly identified modifications that were tolerated without loss of efficacy (+), modifications that resulted in a fivefold loss of efficacy (+/−) and modifications that eliminated efficacy (−). These in vivo results integrate changes in receptor binding and in absorption, distribution, metabolism and excretion (ADME).
One advantage of performing SAR studies in zebrafish is that they couple the analysis of binding affinity and ADME/toxicity. In a traditional in vitro SAR study, structural changes that improve potency are pursued, but these changes might have detrimental effects on absorption, toxicity and so on. By performing SAR studies in whole zebrafish, structures can be identified that improve potency without increased toxicity or loss of in vivo efficacy.
Screens in adult zebrafish
So far, the vast majority of zebrafish experiments have been performed with embryonic and juvenile zebrafish, because the transparency of the fish is greatest at these stages. Furthermore, the embryos and juveniles require much less space than adults and fit easily into multi-well assay plates. A remarkable range of biological and disease process can be studied in these early developmental stages, including disease states such as heart failure and arrhythmias that are frequently associated with adults. However, there are biological processes that are restricted to the adult, and it is becoming clear that the intact adult zebrafish is well-suited to many of these studies. For instance, we have been able to study transplant biology in the zebrafish, having accomplished marrow transplantation in irradiated adults53,54. Studies of regeneration and behaviour have also been carried out in the adult24,55,56. Although the logistics for screening will necessarily be different, it should be possible to carry out chemical screens in whole adult zebrafish.
Toxicology
The rate of innovative pharmaceutical therapies that reach patients seems to be slowing: the number of new molecular entities submitted to the FDA has declined by about half since 1997. In a recent report, the FDA points to technological deficits in toxicology as one of the primary causes of this 'pipeline problem', noting that in many cases, the approaches of the last century are still being used to assess this century's candidates (see online links box). New animal models are needed to test the safety of novel drug candidates, and the FDA report that an estimated 10% improvement in predicting failures before clinical trials would save US $100 million per drug in development costs. In addition to outdated technologies, toxicology frequently suffers by being divorced from the drug discovery process — efforts to discover leads and improve their potency often occur independently from the assessment of toxicity57. Some efforts are being made to involve toxicology earlier in the drug discovery process, such as eliminating compounds with problematic chemical moieties from screening libraries58 or prioritizing leads on the basis of performance in in vitro toxicity assays59. However, much more progress is needed to develop better animal models for toxicological assessment and to involve toxicology earlier in the drug discovery process.
The zebrafish is rapidly gaining acceptance as a promising animal model for toxicology60. The ability to efficiently assess the toxicity of a large number of compounds enables whole libraries to be prescreened for potential toxicity. In this way, compounds with obvious toxicity can be eliminated from libraries before HTS. Alternatively, preliminary hits from HTS can all be tested for toxicity as a means of prioritizing compounds for further development. Significantly, toxicological studies in zebrafish also require compound quantities only in the microgram or milligram ranges, whereas mammalian assays frequently require a few grams to several hundred grams of a given compound.
Most zebrafish toxicity studies so far have focused on environmental contaminants, including pesticides. The zebrafish is not as well established as a model for drug toxicology, and questions remain about how relevant fish toxicity is to humans. Nevertheless, studies have begun to show that many toxic responses are well conserved between fish and mammals. Toxic-response similarities between zebrafish and mammals have been noted for small molecules that cause endocrine disruption, reproductive toxicity, behavioural defects, teratogenesis, carcinogenesis, cardiotoxicity, ototoxicity, liver toxicity and so on61,62,63,64,65,66.
Several zebrafish assays have been developed specifically to monitor toxicities of significance to drug development. For example, Amanuma et al. developed a sensitive zebrafish assay for detecting small-molecule-induced mutagenesis67. Embryos from a zebrafish line that harbour a plasmid containing the 30S ribosomal subunit protein S12 (rpsL) gene can be exposed to a test compound, after which the plasmid is excised from the zebrafish genome and transformed into bacteria. Wild-type copies of the plasmid produce colonies that are resistant to kanamycin but sensitive to streptomycin, whereas mutant copies of the plasmid produce colonies that are resistant to both kanamycin and streptomycin. Therefore, mutation rates can readily be calculated with great sensitivity. Although there are other assays for mutagenesis, including the Ames test and micronucleus test, this assay offers the advantage of testing compounds in a complete organismal context, in which metabolic pathways, DNA repair and other relevant physiological processes are active.
Cardiotoxicity is another area in which a zebrafish assay has the potential to contribute to drug development. Drug-induced prolongation of the cardiac QT interval can lead to the fatal arrhythmia TORSADE DE POINTES, and QT prolongation has become a leading cause of failure during drug development. Milan et al. have developed an automated, high-throughput assay for bradycardia in zebrafish embryos, which they have shown to correlate with QT prolongation in humans68. They tested 100 compounds in the assay and showed that 22 of 23 drugs known to cause QT prolongation in humans cause bradycardia in zebrafish. In addition, the assay was able to detect drug–drug interactions that lead to QT prolongation, such as the well-known synergistic interactions between erythromycin and cisapride and between cimetidine and terfenadine. These interactions result from the physiological effects of one compound influencing the metabolism of the second compound and can only be detected in a whole organism. These results highlight the value of performing toxicity studies in zebrafish — zebrafish assays can achieve the scale and throughput of in vitro assays, but they occur in a relevant physiological setting in which complex pharmacokinetic and pharmacodynamic processes remain intact.
Conclusions
The zebrafish has been pioneered as a developmental and genetic tool for the study of organogenesis and disease. Zebrafish researchers in the basic sciences have taken advantage of the high degree of genetic and physiological similarity between zebrafish and mammals, their small size, large numbers and the fact that phenotypes can be so rapidly assessed in high throughput. These same attributes are beginning to provide opportunities for increases in efficiency in the fields of drug discovery and development. Genetic and morpholino oligonucleotide screens are being used to systematically identify and validate novel therapeutic targets. Zebrafish disease models are being used in whole-organism screens to identify small molecules that suppress the disease state; and zebrafish are proving to be a useful organism for toxicology, allowing the safety of compounds to be assessed earlier, more quickly and less expensively.
What advantages do zebrafish offer over conventional in vitro approaches to target validation, lead discovery and toxicology? First, the whole-organism approach allows biological questions to be addressed that simply cannot be addressed in vitro. For instance, one of our screens has been used to identify small molecules that reverse the cardiac dilation and contractility defects in a zebrafish model of heart failure (C. Hong and R.T.P., unpublished observations). Second, even when an in vitro screen is available, the results obtained by whole-organism screens might be more relevant than those obtained using the in vitro screen. As we have shown in collaboration with the Institute for Chemistry and Cell Biology at Harvard Medical School, many of the small molecules that we identified in our zebrafish cell-cycle screen were not identified in analogous in vitro cell-line screens (L.I.Z., unpublished observations). There could be several reasons for this, including the fact that cancer cell lines with aberrant cell-cycle regulation were used and potential differences between the zebrafish and human cell cycles. However, it is also possible that a physiologically relevant organismal context is required for the mechanisms of action of the compounds that we discovered. Therefore, small molecules identified in whole-organism screens might be more relevant than those identified by in vitro and cell-culture-based screens.
Significant questions remain about how closely zebrafish can model human disease processes and how reliably zebrafish results translate to humans. For example, it is presently unclear how often small molecules discovered in zebrafish screens will have similar effects in humans, nor is it clear exactly how predictive zebrafish toxicity will be of human toxicity. However, initial findings indicate a high degree of similarity between the drug responses of zebrafish and humans69. Our studies of many chemicals that affect the cell cycle show that 50–70% of the chemicals known to affect the cell cycle in mammalian cell culture have similar effects on the zebrafish cell cycle (R. Murphey and L.I.Z, unpublished observations). Other studies indicate an even greater degree of conservation of drug responses, with estimates reaching 95%68. These results might reflect the high degree of amino-acid sequence identity between zebrafish and human drug targets, particularly at protein active sites where many drugs bind. Nonetheless, it is important to realize that zebrafish screening might lead to the identification of small molecules that are species specific, perhaps reflecting differences between zebrafish and human protein structure or ADME.
In recent years, efforts to improve animal studies in drug development have focused on developing 'high-end' animal models that more faithfully mimic human biology70. These efforts have included genetic manipulation of traditional animal models, including rodents, and the adoption of new large animal species. In many instances, large animals and genetically modified small animals have made prediction of drug efficacy and toxicity more reliable. However, as animal models become more sophisticated, they also tend to become more expensive and are necessarily limited to use in a smaller number of experiments. In the near future, the zebrafish is unlikely to contribute to efforts at more faithfully modelling human disease and toxicity. For many applications, primates and other large mammals will continue to be the gold standard for assessing efficacy and safety in the late stages of drug development. However, the zebrafish offers the ability to test the efficacy and safety of thousands of compounds quickly and inexpensively. This approach will not eliminate the need for sophisticated mammalian assays but will be appropriate for many circumstances in which cost, scale and efficiency are more important than perfect replication of human physiology. The rapid, inexpensive prediction of the biological activity of a small molecule might be especially useful for phenotype-based discovery of novel drug leads and in preliminary assessments of compound safety. By providing a low-cost and efficient (albeit imperfect) alternative to other animal models, zebrafish chemical screens possess the hallmarks of a DISRUPTIVE TECHNOLOGY71. Similar to other disruptive technologies, zebrafish chemical screens are likely to allow novel applications that were previously cost-prohibited and to replace other animal models for simple experiments. As technologies improve, the zebrafish might be able to replace mammalian models in applications of increasing complexity.
The zebrafish has already provided a wealth of fundamental information about embryonic development and disease. With the completion of the zebrafish genome project and the establishment of a robust infrastructure for genetic and physiological studies, the zebrafish system sits poised to take on a larger role in the field of drug development. By contributing to target identification and validation, drug lead discovery and toxicology, the zebrafish might provide a shorter path to developing novel therapies for human disease.
References
Eisen, J. S. Zebrafish make a big splash. Cell 87, 969–977 (1996).
Driever, W. et al. A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123, 37–46 (1996).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996). References 2 and 3 describe the first large-scale genetic screens in zebrafish. These screens showed the remarkable potential of zebrafish screens and fuelled the rapid increase in the use of zebrafish for research.
Stickney, H. L. et al. Rapid mapping of zebrafish mutations with SNPs and oligonucleotide microarrays. Genome Res. 12, 1929–1934 (2002).
Ton, C., Stamatiou, D., Dzau, V. J. & Liew, C. C. Construction of a zebrafish cDNA microarray: gene expression profiling of the zebrafish during development. Biochem. Biophys. Res. Commun. 296, 1134–1142 (2002).
Nasevicius, A. & Ekker, S. C. Effective targeted gene 'knockdown' in zebrafish. Nature Genet. 26, 216–220 (2000). This article describes the first use of antisense morpholino oligonucleotides to knock down gene function. Morpholino oligonucleotides continue to be the preferred technique for rapid, reliable gene inactivation in zebrafish.
Heasman, J. Morpholino oligos: making sense of antisense? Dev. Biol. 243, 209–214 (2002).
Meng, A., Jessen, J. R. & Lin, S. Transgenesis. Methods Cell Biol. 60, 133–148 (1999).
Davidson, A. E. et al. Efficient gene delivery and gene expression in zebrafish using the Sleeping Beauty transposon. Dev. Biol. 263, 191–202 (2003).
Kurita, K., Burgess, S. M. & Sakai, N. Transgenic zebrafish produced by retroviral infection of in vitro-cultured sperm. Proc. Natl Acad. Sci. USA 101, 1263–1267 (2004).
Wienholds, E., Schulte-Merker, S., Walderich, B. & Plasterk, R. H. Target-selected inactivation of the zebrafish rag1 gene. Science 297, 99–102 (2002).
Lee, K. Y., Huang, H., Ju, B., Yang, Z. & Lin, S. Cloned zebrafish by nuclear transfer from long-term-cultured cells. Nature Biotechnol. 20, 795–799 (2002).
Bhatt, D. H., Otto, S. J., Depoister, B. & Fetcho, J. R. Cyclic AMP-induced repair of zebrafish spinal circuits. Science 305, 254–258 (2004).
Farber, S. A. et al. Genetic analysis of digestive physiology using fluorescent phospholipid reporters. Science 292, 1385–1388 (2001).
Lindsay, M. A. Target discovery. Nature Rev. Drug Discov. 2, 831–838 (2003).
Bleicher, K. H., Bohm, H. J., Muller, K. & Alanine, A. I. Hit and lead generation: beyond high-throughput screening. Nature Rev. Drug Discov. 2, 369–378 (2003).
Armer, R. E. & Morris, I. D. Trends in early drug safety. Drug News Perspect. 17, 143–148 (2004).
Shin, J. T. & Fishman, M. C. From zebrafish to human: modular medical models. Annu. Rev. Genomics Hum. Genet. 3, 311–340 (2002).
Rubinstein, A. L. Zebrafish: from disease modeling to drug discovery. Curr. Opin. Drug Discov. Devel. 6, 218–223 (2003).
Amatruda, J. F., Shepard, J. L., Stern, H. M. & Zon, L. I. Zebrafish as a cancer model system. Cancer Cell 1, 229–231 (2002).
Amsterdam, A. et al. Identification of 315 genes essential for early zebrafish development. Proc. Natl Acad. Sci. USA 101, 12792–12797 (2004).
Sun, Z. et al. A genetic screen in zebrafish identifies cilia genes as a principal cause of cystic kidney. Development 131, 4085–4093 (2004).
Otto, E. A. et al. Mutations in INVS encoding inversin cause nephronophthisis type 2, linking renal cystic disease to the function of primary cilia and left-right axis determination. Nature Genet. 34, 413–420 (2003).
Poss, K. D., Keating, M. T. & Nechiporuk, A. Tales of regeneration in zebrafish. Dev. Dyn. 226, 202–210 (2003).
Gerull, B. et al. Mutations of TTN, encoding the giant muscle filament titin, cause familial dilated cardiomyopathy. Nature Genet. 30, 201–204 (2002).
Xu, X. et al. Cardiomyopathy in zebrafish due to mutation in an alternatively spliced exon of titin. Nature Genet. 30, 205–209 (2002).
Li, Q. Y. et al. Holt-Oram syndrome is caused by mutations in TBX5, a member of the Brachyury (T) gene family. Nature Genet. 15, 21–29 (1997).
Garrity, D. M., Childs, S. & Fishman, M. C. The heartstrings mutation in zebrafish causes heart/fin Tbx5 deficiency syndrome. Development 129, 4635–4645 (2002).
Curran, M. E. et al. A molecular basis for cardiac arrhythmia: HERG mutations cause long QT syndrome. Cell 80, 795–803 (1995).
Langheinrich, U., Vacun, G. & Wagner, T. Zebrafish embryos express an orthologue of HERG and are sensitive toward a range of QT-prolonging drugs inducing severe arrhythmia. Toxicol. Appl. Pharmacol. 193, 370–382 (2003).
Donovan, A. et al. Positional cloning of zebrafish ferroportin1 identifies a conserved vertebrate iron exporter. Nature 403, 776–781 (2000). This article shows that the hypochromic anaemia phenotype of the zebrafish mutant weissherbst is caused by mutation of ferroportin 1 . This is the first time that a zebrafish gene has led to the discovery of a new human disease gene.
Njajou, O. T. et al. A mutation in SLC11A3 is associated with autosomal dominant hemochromatosis. Nature Genet. 28, 213–214 (2001).
Davidson, A. J. et al. cdx4 mutants fail to specify blood progenitors and can be rescued by multiple hox genes. Nature 425, 300–306 (2003).
Kalev-Zylinska, M. L. et al. Runx1 is required for zebrafish blood and vessel development and expression of a human RUNX1-CBF2T1 transgene advances a model for studies of leukemogenesis. Development 129, 2015–2030 (2002).
Langenau, D. M. et al. Myc-induced T cell leukemia in transgenic zebrafish. Science 299, 887–890 (2003).
Li, L. & Dowling, J. E. A dominant form of inherited retinal degeneration caused by a non-photoreceptor cell-specific mutation. Proc. Natl Acad. Sci. USA 94, 11645–11650 (1997).
Peitsaro, N., Kaslin, J., Anichtchik, O. V. & Panula, P. Modulation of the histaminergic system and behaviour by α-fluoromethylhistidine in zebrafish. J. Neurochem. 86, 432–441 (2003).
Nicolson, T. et al. Genetic analysis of vertebrate sensory hair cell mechanosensation: the zebrafish circler mutants. Neuron 20, 271–283 (1998).
Malicki, J. et al. Mutations affecting development of the zebrafish ear. Development 123, 275–283 (1996).
Neuhauss, S. C. et al. Genetic disorders of vision revealed by a behavioral screen of 400 essential loci in zebrafish. J. Neurosci. 19, 8603–8615 (1999).
Darland, T. & Dowling, J. E. Behavioral screening for cocaine sensitivity in mutagenized zebrafish. Proc. Natl Acad. Sci. USA 98, 11691–11696 (2001).
Carpinelli, M. R. et al. Suppressor screen in Mpl−/− mice: c-Myb mutation causes supraphysiological production of platelets in the absence of thrombopoietin signaling. Proc. Natl Acad. Sci. USA 101, 6553–6558 (2004). In this study, a genetic screen identified mutations in c-Myb that can suppress the disease phenotype in a murine model of thrombocytopaenia. This article shows the feasibility of using genetic screens to identify novel drug targets.
Kenwrick, S., Amaya, E. & Papalopulu, N. Pilot morpholino screen in Xenopus tropicalis identifies a novel gene involved in head development. Dev. Dyn. 229, 289–299 (2004).
Yamada, L. et al. Morpholino-based gene knockdown screen of novel genes with developmental function in Ciona intestinalis. Development 130, 6485–6495 (2003).
Tobert, J. A. Lovastatin and beyond: the history of the HMG-CoA reductase inhibitors. Nature Rev. Drug Discov. 2, 517–526 (2003).
Clader, J. W. The discovery of ezetimibe: a view from outside the receptor. J. Med. Chem. 47, 1–9 (2004).
Altmann, S. W. et al. Niemann-Pick C1 Like 1 protein is critical for intestinal cholesterol absorption. Science 303, 1201–1204 (2004).
Robson, J. M. & Keele, C. A Recent Advances in Pharmacology (J. & A. Churchill, London 1956).
Peterson, R. T. et al. Chemical suppression of a genetic mutation in a zebrafish model of aortic coarctation. Nature Biotechnol. 22, 595–599 (2004). This article describes the discovery of compounds that completely suppress the phenotypic effects of a zebrafish vascular mutation, revealing the potential of zebrafish screens for discovering disease-suppressing compounds.
Stern, H. M. & Zon, L. I. Cancer genetics and drug discovery in the zebrafish. Nature Rev. Cancer 3, 533–539 (2003).
Zhong, T. P., Rosenberg, M., Mohideen, M. A., Weinstein, B. & Fishman, M. C. gridlock, an HLH gene required for assembly of the aorta in zebrafish. Science 287, 1820–1824 (2000).
Zhong, T. P., Childs, S., Leu, J. P. & Fishman, M. C. Gridlock signalling pathway fashions the first embryonic artery. Nature 414, 216–220 (2001).
Langenau, D. M. et al. In vivo tracking of T cell development, ablation, and engraftment in transgenic zebrafish. Proc. Natl Acad. Sci. USA 101, 7369–7374 (2004).
Traver, D. et al. Effects of lethal irradiation in zebrafish and rescue by hematopoietic cell transplantation. Blood 104, 1298–1305 (2004).
Poss, K. D., Wilson, L. G. & Keating, M. T. Heart regeneration in zebrafish. Science 298, 2188–2190 (2002).
Guo, S. Linking genes to brain, behavior and neurological diseases: what can we learn from zebrafish? Genes Brain Behav. 3, 63–74 (2004).
Ulrich, R. & Friend, S. H. Toxicogenomics and drug discovery: will new technologies help us produce better drugs? Nature Rev. Drug Discov. 1, 84–88 (2002).
Llorens, O., Perez, J. J. & Villar, H. O. Toward the design of chemical libraries for mass screening biased against mutagenic compounds. J. Med. Chem. 44, 2793–2804 (2001).
Pritchard, J. F. et al. Making better drugs: decision gates in non-clinical drug development. Nature Rev. Drug Discov. 2, 542–553 (2003).
Spitsbergen, J. M. & Kent, M. L. The state of the art of the zebrafish model for toxicology and toxicologic pathology research — advantages and current limitations. Toxicol. Pathol. 31 Suppl, 62–87 (2003).
Parng, C., Seng, W. L., Semino, C. & McGrath, P. Zebrafish: a preclinical model for drug screening. Assay Drug Dev. Technol. 1, 41–48 (2002).
Brion, F. et al. Impacts of 17β-estradiol, including environmentally relevant concentrations, on reproduction after exposure during embryo-larval-, juvenile- and adult-life stages in zebrafish (Danio rerio). Aquat. Toxicol. 68, 193–217 (2004).
Levin, E. D., Chrysanthis, E., Yacisin, K. & Linney, E. Chlorpyrifos exposure of developing zebrafish: effects on survival and long-term effects on response latency and spatial discrimination. Neurotoxicol. Teratol. 25, 51–57 (2003).
Schilling, T. F. & Knight, R. D. Origins of anteroposterior patterning and Hox gene regulation during chordate evolution. Philos. Trans. R Soc. Lond. B Biol. Sci. 356, 1599–1613 (2001).
Spitsbergen, J. M. et al. Neoplasia in zebrafish (Danio rerio) treated with 7,12-dimethylbenz[a]anthracene by two exposure routes at different developmental stages. Toxicol. Pathol. 28, 705–715 (2000).
Murakami, S. L. et al. Developmental differences in susceptibility to neomycin-induced hair cell death in the lateral line neuromasts of zebrafish (Danio rerio). Hear. Res. 186, 47–56 (2003).
Amanuma, K., Takeda, H., Amanuma, H. & Aoki, Y. Transgenic zebrafish for detecting mutations caused by compounds in aquatic environments. Nature Biotechnol. 18, 62–65 (2000).
Milan, D. J., Peterson, T. A., Ruskin, J. N., Peterson, R. T. & MacRae, C. A. Drugs that induce repolarization abnormalities cause bradycardia in zebrafish. Circulation 107, 1355–1358 (2003). In this study, the cardiotoxicity of 100 compounds was tested using an automated zebrafish assay. A high degree of correlation was observed between zebrafish toxicity and known human toxicities, indicating that zebrafish might be well suited for studies of cardiotoxicity.
Langheinrich, U. Zebrafish: a new model on the pharmaceutical catwalk. Bioessays 25, 904–912 (2003).
Greaves, P., Williams, A. & Eve, M. First dose of potential new medicines to humans: how animals help. Nature Rev. Drug Discov. 3, 226–236 (2004).
Christensen, C. M. The Innovator's Dilemma: When New Technologies Cause Great Firms to Fail (Harvard Business School Press, Boston, Massachusetts, 1997).
Solnica-Krezel, L., Schier, A. F. & Driever, W. Efficient recovery of ENU-induced mutations from the zebrafish germline. Genetics 136, 1401–1420 (1994).
Amsterdam, A. Insertional mutagenesis in zebrafish. Dev. Dyn. 228, 523–534 (2003).
Kwok, C. et al. Characterization of whole genome radiation hybrid mapping resources for non-mammalian vertebrates. Nucleic Acids Res. 26, 3562–3566 (1998).
Hukriede, N. A. et al. Radiation hybrid mapping of the zebrafish genome. Proc. Natl Acad. Sci. USA 96, 9745–9750 (1999).
Geisler, R. et al. A radiation hybrid map of the zebrafish genome. Nature Genet. 23, 86–89 (1999).
Affymetrix GeneChip Array Products [online], <http://www.affymetrix.com/products/arrays/>
Zebrafish Genome Resources [online], <http://www.ncbi.nlm.nih.gov/genome/guide/zebrafish/>
Zebrafish International Resource Center [online], <http://zfin.org/zirc/home/guide.php>
Tübingen Zebrafish Stockcenter [online], <http://www.eb.tuebingen.mpg.de/services/stockcenter/>
The Sanger Institute Danio rerio Sequencing Project [online], <http://www.sanger.ac.uk/Projects/D_rerio/>
Trans-NIH Zebrafish Initiative [online], <http://www.nih.gov/science/models/zebrafish/>
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
R.T.P. receives research funding through a sponsored research agreement with Novartis Institutes for BioMedical Research.
Related links
Related links
DATABASES
Entrez Gene
OMIM
FURTHER INFORMATION
Glossary
- ORGANOGENESIS
-
The formation of organs during ontogeny.
- MORPHOLINO OLIGONUCLEOTIDE
-
A synthetic oligonucleotide with a backbone that has been chemically modified to resist nucleotide degradation.
- MAUTHNER CELLS
-
A bilateral pair of brain stem neurons that receive sensory inputs and trigger the characteristic escape response of fish to aversive stimuli.
- HAEMOCHROMATOSIS TYPE 4
-
A hereditary condition that is characterized by the abnormal accumulation of iron in tissues of the body.
- CONDITIONED PLACE PREFERENCE
-
An experimental procedure to test an animal's learned association between environmental cues and the subjective state produced by a drug. After repeated exposure to a drug in the presence of distinctive environmental cues, the animal is tested to see if it will preferentially move towards a compartment that contains drug-associated environmental cues, even in the absence of the drug.
- THROMBOCYTOPAENIA
-
A condition characterized by an abnormal reduction in the number of circulating platelets.
- TORSADE DE POINTES
-
A form of polymorphous ventricular tachycardia that is associated with prolongation of the cardiac QT interval that can lead to sudden cardiac death.
- DISRUPTIVE TECHNOLOGY
-
A new technology that is simpler and cheaper than the prevailing technology but initially offers reduced performance. As the technology's capabilities improve, its simplicity and cost might allow it to supplant the prevailing technology.
Rights and permissions
About this article
Cite this article
Zon, L., Peterson, R. In vivo drug discovery in the zebrafish. Nat Rev Drug Discov 4, 35–44 (2005). https://doi.org/10.1038/nrd1606
Issue Date:
DOI: https://doi.org/10.1038/nrd1606
This article is cited by
-
Preclinical models in head and neck squamous cell carcinoma
British Journal of Cancer (2023)
-
Biodistribution and toxicity assessment of methoxyphenyl phosphonium carbosilane dendrimers in 2D and 3D cell cultures of human cancer cells and zebrafish embryos
Scientific Reports (2023)
-
Construction of multiple concentration gradients for single-cell level drug screening
Microsystems & Nanoengineering (2023)
-
Closing the loop between brain and electrical stimulation: towards precision neuromodulation treatments
Translational Psychiatry (2023)
-
Near-infrared-II deep tissue fluorescence microscopy and application
Nano Research (2023)