Key Points
-
cyclic AMP and cyclic GMP are universal second messengers and are sensed by receptor proteins that contain cyclic nucleotide binding (CNB) domains.
-
In mammal, CNB domains are found in protein kinase A (PKA), protein kinase G (PKG), the guanine nucleotide-exchange factor Epac and cyclic-nucleotide-regulated ion channels. These receptor proteins mediate the wide range of biological effects that are controlled by cyclic nucleotides, including the regulation of gene transcription, cell adhesion and heart rhythm, mediating the biochemical processes that are fundamental to vision and scent, and the modulation of insulin secretion.
-
CNB domains are structurally conserved switch domains that consist of a central β-sandwich flanked at the N terminus by the N-terminal helical bundle and at the C terminus by the hinge helix and the lid.
-
Ligand binding induces conformational changes in that region of the β-sandwich that interacts directly with the phosphate-sugar moiety of the nucleotide. These changes are translated into a movement of the hinge and the lid, as well as of a rearrangement of the N-terminal helical bundle.
-
The structural changes that are induced by ligand binding are universal in CNB domains. They are coupled in an individual manner with protein activation in the different classes of cyclic nucleotide receptors.
Abstract
Fifty years ago, cyclic AMP was discovered as a second messenger of hormone action, heralding the age of signal transduction. Many cellular processes were found to be regulated by cAMP and the related cyclic GMP. Cyclic nucleotides function by binding to and activating their effectors — protein kinase A, protein kinase G, cyclic-nucleotide-regulated ion channels and the guanine nucleotide-exchange factor Epac. Recent structural insights have now made it possible to propose a general structural mechanism for how cyclic nucleotides regulate these proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Vaandrager, A. B. & de Jonge, H. R. Signalling by cGMP-dependent protein kinases. Mol. Cell. Biochem. 157, 23–30 (1996).
Craven, K. B. & Zagotta, W. N. CNG and HCN channels: two peas, one pod. Annu. Rev. Physiol. 68, 375–401 (2006).
Kaupp, U. B. & Seifert, R. Molecular diversity of pacemaker ion channels. Annu. Rev. Physiol. 63, 235–257 (2001).
Kaupp, U. B. & Seifert, R. Cyclic nucleotide-gated ion channels. Physiol. Rev. 82, 769–824 (2002).
Bos, J. L. Epac: a new cAMP target and new avenues in cAMP research. Nature Rev. Mol. Cell Biol. 4, 733–738 (2003).
Seino, S. & Shibasaki, T. PKA-dependent and PKA-independent pathways for cAMP-regulated exocytosis. Physiol. Rev. 85, 1303–1342 (2005).
Beavo, J. A. & Brunton, L. L. Cyclic nucleotide research — still expanding after half a century. Nature Rev. Mol. Cell Biol. 3, 710–718 (2002).
Rall, T. W., Sutherland, E. W. & Berthet, J. The relationship of epinephrine and glucagon to liver phosphorylase. IV. Effect of epinephrine and glucagon on the reactivation of phosphorylase in liver homogenates. J. Biol. Chem. 224, 463–475 (1957).
Sutherland, E. W. & Rall, T. W. The properties of an adenine ribonucleotide produced with cellular particles, ATP, Mg++, and epinephrine or glucagon. J. Am. Chem. Soc. 79, 3608 (1957).
Cook, W. H., Lipkin, D. & Markham, R. The formation of a cyclic dianhydrodiadenylic acid (I) by the alkaline degradation of adenosine-5′-triphosphoric acid (II). J. Am. Chem. Soc. 79, 3607–3608 (1957).
Lipkin, D., Cook, W. H. & Markham, R. Adenosine-3′:5′-phosphoric acid: a proof of structure. J. Am. Chem. Soc. 81, 6198–6203 (1959).
Sutherland, E. W., Rall, T. W. & Menon, T. Adenyl cyclase. I. Distribution, preperation, and properties. J. Biol. Chem. 237, 1220 (1961).
Taussig, R. & Gilman, A. G. Mammalian membrane-bound adenylyl cyclases. J. Biol. Chem. 270, 1–4 (1995).
Hanoune, J. & Defer, N. Regulation and role of adenylyl cyclase isoforms. Annu. Rev. Pharmacol. Toxicol. 41, 145–174 (2001).
Taussig, R. & Zimmermann, G. Type-specific regulation of mammalian adenylyl cyclases by G protein pathways. Adv. Sec. Mess. Phos. Res. 32, 81–98 (1998).
Walsh, D. A., Perkins, J. P. & Krebs, E. G. Adensine 3′,5′-monophosphate-dependant protein kinase from rabbit skeletal muscle. J. Biol. Chem. 243, 3763–3765 (1968).
Ashman, D. F., Lipton, R., Melicow, M. M. & Price, T. D. Isolation of adenosine 3′,5′-monophosphate and guanosine 3′,5′-monophosphate from rat urine. Biochem. Biophys. Res. Commun. 11, 330–334 (1963).
Kuo, J. F. & Greengard, P. Cyclic nucleotide-dependent protein kinases. IV. Widespread occurrence of adenosine 3′,5′-monophosphate-dependent protein kinase in various tissues and phyla of the animal kingdom. Proc. Natl Acad. Sci. USA 64, 1349–1355 (1969).
Kuo, J. F. & Greengard, P. Cyclic nucleotide-dependent protein kinases. VI. Isolation and partial purification of a protein kinase activated by guanosine 3′,5′-monophosphate. J. Biol. Chem. 245, 2493–2498 (1970).
Yuen, P. S. & Garbers, D. L. Guanylyl cyclase-linked receptors. Annu. Rev. Neurosci. 15, 193–225 (1992).
Currie, M. G. et al. Bioactive cardiac substances: potent vasorelaxant activity in mammalian atria. Science 221, 71–73 (1983).
Furchgott, R. F. & Zawadzki, J. V. The obligatory role of endothelial cells in the relaxation of arterial smooth muscle by acetylcholine. Nature 288, 373–376 (1980).
Palmer, R. M., Ferrige, A. G. & Moncada, S. Nitric oxide release accounts for the biological activity of endothelium-derived relaxing factor. Nature 327, 524–526 (1987).
Ignarro, L. J., Buga, G. M., Wood, K. S., Byrns, R. E. & Chaudhuri, G. Endothelium-derived relaxing factor produced and released from artery and vein is nitric oxide. Proc. Natl Acad. Sci. USA 84, 9265–9269 (1987).
Friebe, A. & Koesling, D. Regulation of nitric oxide-sensitive guanylyl cyclase. Circ. Res. 93, 96–105 (2003).
Beavo, J. A., Conti, M. & Heaslip, R. J. Multiple cyclic nucleotide phosphodiesterases. Mol. Pharmacol. 46, 399–405 (1994).
Manganiello, V. C., Murata, T., Taira, M., Belfrage, P. & Degerman, E. Diversity in cyclic nucleotide phosphodiesterase isoenzyme families. Arch. Biochem. Biophys. 322, 1–13 (1995).
Aravind, L. & Ponting, C. P. The GAF domain: an evolutionary link between diverse phototransducing proteins. Trends Biochem. Sci. 22, 458–459 (1997).
Martinez, S. E., Beavo, J. A. & Hol, W. G. GAF domains: two-billion-year-old molecular switches that bind cyclic nucleotides. Mol. Interv. 2, 317–323 (2002).
Zoraghi, R., Corbin, J. D. & Francis, S. H. Properties and functions of GAF domains in cyclic nucleotide phosphodiesterases and other proteins. Mol. Pharmacol. 65, 267–278 (2004).
Gross-Langenhoff, M., Hofbauer, K., Weber, J., Schultz, A. & Schultz, J. E. cAMP is a ligand for the tandem GAF domain of human phosphodiesterase 10 and cGMP for the tandem GAF domain of phosphodiesterase 11. J. Biol. Chem. 281, 2841–2846 (2006).
Martinez, S. E. et al. The two GAF domains in phosphodiesterase 2A have distinct roles in dimerization and in cGMP binding. Proc. Natl Acad. Sci. USA 99, 13260–13265 (2002).
Martinez, S. E. et al. Crystal structure of the tandem GAF domains from a cyanobacterial adenylyl cyclase: modes of ligand binding and dimerization. Proc. Natl Acad. Sci. USA 102, 3082–3087 (2005).
Fesenko, E. E., Kolesnikov, S. S. & Lyubarsky, A. L. Induction by cyclic GMP of cationic conductance in plasma membrane of retinal rod outer segment. Nature 313, 310–313 (1985).
Miki, N., Keirns, J. J., Marcus, F. R., Freeman, J. & Bitensky, M. W. Regulation of cyclic nucleotide concentrations in photoreceptors: an ATP-dependent stimulation of cyclic nucleotide phosphodiesterase by light. Proc. Natl Acad. Sci. USA 70, 3820–3824 (1973).
Haynes, L. & Yau, K. W. Cyclic GMP-sensitive conductance in outer segment membrane of catfish cones. Nature 317, 61–64 (1985).
Cook, N. J., Hanke, W. & Kaupp, U. B. Identification, purification, and functional reconstitution of the cyclic GMP-dependent channel from rod photoreceptors. Proc. Natl Acad. Sci. USA 84, 585–589 (1987).
Nakamura, T. & Gold, G. H. A cyclic nucleotide-gated conductance in olfactory receptor cilia. Nature 325, 442–444 (1987).
Kaupp, U. B. et al. Primary structure and functional expression from complementary DNA of the rod photoreceptor cyclic GMP-gated channel. Nature 342, 762–766 (1989).
de Rooij, J. et al. Epac is a Rap1 guanine-nucleotide-exchange factor directly activated by cyclic AMP. Nature 396, 474–477 (1998).
Kawasaki, H. et al. A family of cAMP-binding proteins that directly activate Rap1. Science 282, 2275–2279 (1998).
McKay, D. B. & Steitz, T. A. Structure of catabolite gene activator protein at 2. 9 Å resolution suggests binding to left-handed B-DNA. Nature 290, 744–749 (1981). The first structure of a CNB domain; that of the bacterial protein CAP.
Emmer, M., deCrombrugghe, B., Pastan, I. & Perlman, R. Cyclic AMP receptor protein of E. coli: its role in the synthesis of inducible enzymes. Proc. Natl Acad. Sci. USA 66, 480–487 (1970).
Kolb, A., Busby, S., Buc, H., Garges, S. & Adhya, S. Transcriptional regulation by cAMP and its receptor protein. Annu. Rev. Biochem. 62, 749–795 (1993).
Bell, A. et al. Mutations that alter the ability of the Escherichia coli cyclic AMP receptor protein to activate transcription. Nucleic Acids Res. 18, 7243–7250 (1990).
Aiba, H., Fujimoto, S. & Ozaki, N. Molecular cloning and nucleotide sequencing of the gene for E. coli cAMP receptor protein. Nucleic Acids Res. 10, 1345–1361 (1982).
Cossart, P. & Gicquel-Sanzey, B. Cloning and sequence of the crp gene of Escherichia coli K12. Nucleic Acids Res. 10, 1363–1378 (1982).
McKay, D. B., Weber, I. T. & Steitz, T. A. Structure of catabolite gene activator protein at 2.9-Å resolution. Incorporation of amino acid sequence and interactions with cyclic AMP. J. Biol. Chem. 257, 9518–9524 (1982).
Takio, K., Smith, S. B., Krebs, E. G., Walsh, K. A. & Titani, K. Primary structure of the regulatory subunit of type II cAMP-dependent protein kinase from bovine cardiac muscle. Proc. Natl Acad. Sci. USA 79, 2544–2548 (1982).
Titani, K. et al. Amino acid sequence of the regulatory subunit of bovine type I adenosine cyclic 3′,5′-phosphate dependent protein kinase. Biochemistry 23, 4193–4199 (1984).
Takio, K. et al. Guanosine cyclic 3′,5′-phosphate dependent protein kinase, a chimeric protein homologous with two separate protein families. Biochemistry 23, 4207–4218 (1984).
Weber, I. T., Takio, K., Titani, K. & Steitz, T. A. The cAMP-binding domains of the regulatory subunit of cAMP-dependent protein kinase and the catabolite gene activator protein are homologous. Proc. Natl Acad. Sci. USA 79, 7679–7683 (1982).
Corbin, J. D. et al. Studies on the properties and mode of action of the purified regulatory subunit of bovine heart adenosine 3′:5′-monophosphate-dependent protein kinase. J. Biol. Chem. 253, 3997–4003 (1978).
Su, Y. et al. Regulatory subunit of protein kinase A: structure of deletion mutant with cAMP binding domains. Science 269, 807–813 (1995).
Diller, T. C., Madhusudan, Xuong, N. H. & Taylor, S. S. Molecular basis for regulatory subunit diversity in cAMP-dependent protein kinase: crystal structure of the type II β regulatory subunit. Structure 9, 73–82 (2001). References 54 and 55 describe the structure of the regulatory subunit type I and type II, respectively.
Kim, C., Xuong, N. H. & Taylor, S. S. Crystal structure of a complex between the catalytic and regulatory (RIα) subunits of PKA. Science 307, 690–696 (2005). Structure of the complex between the regulatory subunit and the catalytic subunit of PKA.
Rehmann, H. et al. Structure and regulation of the cAMP-binding domains of Epac2. Nature Struct. Biol. 10, 26–32 (2003). First structure of a CNB domain in the absence of cyclic nucleotide; the hinge model was first presented here.
Rehmann, H., Das, J., Knipscheer, P., Wittinghofer, A. & Bos, J. L. Structure of the cyclic-AMP-responsive exchange factor Epac2 in its auto-inhibited state. Nature 439, 625–628 (2006).
Zagotta, W. N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003).
Clayton, G. M., Silverman, W. R., Heginbotham, L. & Morais-Cabral, J. H. Structural basis of ligand activation in a cyclic nucleotide regulated potassium channel. Cell 119, 615–627 (2004). References 59 and 60 describe the structures of CNB domains from cyclic-nucleotide-regulated ion channels.
Shabb, J. B. & Corbin, J. D. Cyclic nucleotide-binding domains in proteins having diverse functions. J. Biol. Chem. 267, 5723–5726 (1992).
Canaves, J. M. & Taylor, S. S. Classification and phylogenetic analysis of the cAMP-dependent protein kinase regulatory subunit family. J. Mol. Evol. 54, 17–19 (2002).
Rehmann, H., Schwede, F., Doskeland, S. O., Wittinghofer, A. & Bos, J. L. Ligand-mediated activation of the cAMP-responsive guanine nucleotide exchange factor Epac. J. Biol. Chem. 278, 38548–38556 (2003).
Enserink, J. M. et al. A novel Epac-specific cAMP analogue demonstrates independent regulation of Rap1 and ERK. Nature Cell Biol. 4, 901–906 (2002).
Weber, I. T., Shabb, J. B. & Corbin, J. D. Predicted structures of the cGMP binding domains of the cGMP-dependent protein kinase: a key alanine/threonine difference in evolutionary divergence of cAMP and cGMP binding sites. Biochemistry 28, 6122–6127 (1989).
Weber, I. T. & Steitz, T. A. Structure of a complex of catabolite gene activator protein and cyclic AMP refined at 2.5 Å resolution. J. Mol. Biol. 198, 311–326 (1987).
Berman, H. M. et al. The cAMP binding domain: an ancient signaling module. Proc. Natl Acad. Sci. USA 102, 45–50 (2005).
Rehmann, H., Rueppel, A., Bos, J. L. & Wittinghofer, A. Communication between the regulatory and the catalytic region of the cAMP-responsive guanine nucleotide exchange factor Epac. J. Biol. Chem. 278, 23508–23514 (2003).
Beavo, J. A., Bechtel, P. J. & Krebs, E. G. Mechanisms of control for cAMP-dependent protein kinase from skeletal muscle. Adv. Cyclic Nucleotide Res. 5, 241–251 (1975).
Saraswat, L. D., Ringheim, G. E., Bubis, J. & Taylor, S. S. Deletion mutants as probes for localizing regions of subunit interaction in cAMP-dependent protein kinase. J. Biol. Chem. 263, 18241–18246 (1988).
Knighton, D. R. et al. Structure of a peptide inhibitor bound to the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 414–420 (1991).
Knighton, D. R. et al. Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 407–414 (1991). References 71 and 72 describe the structure of the catalytic subunit of PKA, which represents the first structure of a kinase domain.
Feramisco, J. R. & Krebs, E. G. Inhibition of cyclic AMP-dependent protein kinase by analogues of a synthetic peptide substrate. J. Biol. Chem. 253, 8968–8971 (1978).
Hofmann, F. Apparent constants for the interaction of regulatory and catalytic subunit of cAMP-dependent protein kinase I and II. J. Biol. Chem. 255, 1559–1564 (1980).
Anand, G. S., Hughes, C. A., Jones, J. M., Taylor, S. S. & Komives, E. A. Amide H/2H exchange reveals communication between the cAMP and catalytic subunit-binding sites in the R(I)α subunit of protein kinase A. J. Mol. Biol. 323, 377–386 (2002).
Hamuro, Y. et al. Dynamics of cAPK type IIβ activation revealed by enhanced amide H/2H exchange mass spectrometry (DXMS). J. Mol. Biol. 327, 1065–1076 (2003).
Zhao, J., Hoye, E., Boylan, S., Walsh, D. A. & Trewhella, J. Quaternary structures of a catalytic subunit-regulatory subunit dimeric complex and the holoenzyme of the cAMP-dependent protein kinase by neutron contrast variation. J. Biol. Chem. 273, 30448–30459 (1998).
Vigil, D., Blumenthal, D. K., Taylor, S. S. & Trewhella, J. The conformationally dynamic C helix of the RIα subunit of protein kinase A mediates isoform-specific domain reorganization upon C subunit binding. J. Biol. Chem. 280, 35521–35527 (2005).
Wolfe, L., Francis, S. H. & Corbin, J. D. Properties of a cGMP-dependent monomeric protein kinase from bovine aorta. J. Biol. Chem. 264, 4157–4162 (1989).
Aggarwal, S. K. & MacKinnon, R. Contribution of the S4 segment to gating charge in the Shaker K+ channel. Neuron 16, 1169–1177 (1996).
Alioua, A. et al. The large conductance, voltage-dependent, and calcium-sensitive K+ channel, Hslo, is a target of cGMP-dependent protein kinase phosphorylation in vivo. J. Biol. Chem. 273, 32950–32956 (1998).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Kuo, A. et al. Crystal structure of the potassium channel KirBac1.1 in the closed state. Science 300, 1922–1926 (2003).
Jiang, Y. et al. The open pore conformation of potassium channels. Nature 417, 523–526 (2002).
Craven, K. B. & Zagotta, W. N. Salt bridges and gating in the COOH-terminal region of HCN2 and CNGA1 channels. J. Gen. Physiol. 124, 663–677 (2004).
Johnson, J. P. Jr. & Zagotta, W. N. The carboxyl-terminal region of cyclic nucleotide-modulated channels is a gating ring, not a permeation path. Proc. Natl Acad. Sci. USA 102, 2742–2747 (2005).
Wainger, B. J., De Gennaro, M., Santoro, B., Siegelbaum, S. A. & Tibbs, G. R. Molecular mechanism of cAMP modulation of HCN pacemaker channels. Nature 411, 805–810 (2001).
de Rooij, J. et al. Mechanism of regulation of the Epac family of cAMP-dependent RapGEFs. J. Biol. Chem. 275, 20829–20836 (2000).
Herrmann, C. Ras-effector interactions: after one decade. Curr. Opin. Struct. Biol. 13, 122–129 (2003).
Coccetti, P., Mauri, I., Alberghina, L., Martegani, E. & Parmeggiani, A. The minimal active domain of the mouse ras exchange factor CDC25Mm. Biochem. Biophys. Res. Commun. 206, 253–259 (1995).
Boriack-Sjodin, P. A., Margarit, S. M., Bar-Sagi, D. & Kuriyan, J. The structural basis of the activation of Ras by Sos. Nature 394, 337–343 (1998).
Botelho, L. H., Rothermel, J. D., Coombs, R. V. & Jastorff, B. cAMP analog antagonists of cAMP action. Methods Enzymol. 159, 159–172 (1988).
Wu, J., Jones, J. M., Xuong, N. H., Ten Eyck, L. F. & Taylor, S. S. Crystal structures of RIα subunit of cyclic adenosine 5′-monophosphate (cAMP)-dependent protein kinase complexed with (Rp)-adenosine 3′,5′-cyclic monophosphothioate and (Sp)-adenosine 3′,5′-cyclic monophosphothioate, the phosphothioate analogues of cAMP. Biochemistry 43, 6620–6629 (2004).
Kraulis, P. J. Molscript — a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E. A. & Murphy, M. E. P. Rasert3D version-2.0 — a program for photorealistic molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 50, 869–873 (1994).
Acknowledgements
We thank F. Zwartkruis and B. Burgering for stimulating discussions. H.R. is supported by the Chemical Sciences of the Netherlands Organization for Scientific Research (NWO-CW).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (movie)
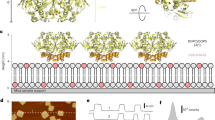
Cyclic-nucleotide-induced conformational changes in cyclic-nucleotide-binding domains. A schematic representation of a cyclic-nucleotide-binding domain is shown. The central core of the domain (the β- sandwich) is depicted in yellow and grey and being barrel shaped. The N-terminal helical bundle, the hinge and the lid are represented as orange helices and labelled correspondingly. The movie shows an animation of the conformational changes that occur in cyclic-nucleotide-binding domains following the binding of cyclic nucleotide (see also FIG. 4). The cyclic nucleotide (shown in red with the phosphate group (P) coloured blue) enters the binding site, and the phosphate-binding cassette (PBC) undergoes rearrangements to allow it to interact with the phosphate sugar moiety of the cyclic nucleotide (red dotted lines represent hydrogen bonds). As a result, the Leu residue at position six in the PBC, which is shown in a red stick representation, is repositioned and its movement leaves a space that allows a conserved Phe or Tyr residue (a Phe is shown here in orange) in the hinge to move closer to the core of the domain (the β-sandwich). A rearrangement of the N-terminal helical bundle is associated with the movement of the hinge. When the hinge adopts its final conformation, it positions the lid over the cyclic nucleotide and the conformation of the lid is stabilized by mainly hydrophobic interactions with the base of the nucleotide (black and red striped lines represent hydrophobic interactions). (MOV 1960 kb)
Glossary
- Glycogen
-
A polysaccharide that is composed of chains of glucose molecules. Glycogen is a storage form of glucose in the body.
- Heterotrimeric G-protein
-
Comprises an α-, β- and γ-subunit. The α-subunit binds GDP or GTP. Exchange of GDP to GTP is stimulated by agonist-stimulated transmembrane receptors and causes dissociation into the α-subunit and a βγ-complex, which control downstream signalling cascades.
- Natriuresis
-
The excretion of an excessively large amount of Na+ ions in the urine. The consequence of natriuresis is that extracellular liquid volume and, therefore, blood pressure are reduced.
- Atrial natriuretic peptide
-
(ANP). A small protein that is mainly released by endocrine heart cells following distension of the right atrium. ANP is generated by the cleavage of its precursor protein.
- GAF domain
-
Named GAF because it is found in cGMP-specific and cGMP-stimulated phosphodiesterases, Anabaena adenylyl cyclases and Escherichia coli FhlA proteins. When present in phosphodiesterases or adenylyl cyclases, this domain usually binds cGMP and cAMP. In some plant and cyanobacterial phytochromes, a GAF domain is the site of chromophore attachment.
- HCN channel
-
(Hyperpolarization-activated, cyclic-nucleotide-modulated channel). A cationic ion channel that is regulated mainly by the membrane potential. Cyclic nucleotide binding modifies the activation characteristics of the channel by shifting the threshold membrane potential needed for channel opening.
- Lac operon
-
A group of three genes that are involved in the metabolism of lactose. The expression of these genes is controlled by a single promoter that is activated by catabolite activator protein (CAP).
- PKA type II (and type I)
-
In higher organisms, four different regulatory subunits of PKA, namely type Iα, type Iβ, type IIα and type IIβ, exist. PKAs can be stimulated by cAMP and can then phosphorylate downstream targets.
- Leucine zipper
-
A specialized coiled-coil motif that can induce the dimerization of α-helices. The helices are held together by hydrophobic interactions between the Leu residues that are located on one side of the amphipathic helices.
- Pseudo-substrate sequence
-
A sequence that resembles the consensus sequence of a kinase, with the exception that the residue that normally accepts the γ-phosphate group is replaced by an Ala residue. The pseudo-substrate sequence binds to the catalytic cleft of the kinase domain and therefore traps the catalytic subunit in an unproductive complex.
- Deuterium-exchange method
-
A technique to measure the accessibility of residues to solvent. Time-limited exposure of a protein to deuterium oxide (heavy water) results in the exchange of solvent-exposed protons by deuterium, which can be analysed by mass spectrometry.
- CNG channel
-
(Cyclic-nucleotide-gated channel). The binding of cyclic nucleotides to this cationic ion channel directly controls the probability of the channel being open. Membrane potential has almost no effect on the regulation of CNG channels.
- Gating current
-
The current that is measured during channel opening and that is independent of the ion flux across the membrane. The current originates from the movement of charged amino-acid residues due to the conformational changes that occur in the channel during opening.
- DEP domain
-
(Dishevelled, Egl-10, Pleckstrin domain). A 90amino-acid domain that is thought to be involved in the membrane localization of the protein in which the domain is present, although the exact targeting mechanism is unknown.
- REM domain
-
(Ras-exchanger-motif domain). A domain that is present in guanine nucleotide-exchange factors for small G-proteins of the Ras family. They mainly provide structural shielding and stabilizing roles.
- RA domain
-
(Ras-association domain; RalGDS/AF6 domain). A domain with a ubiquitin-like fold that is found in effectors of G-proteins of the Ras superfamily. RA domains interact specifically with the GTP-bound conformation of these G-proteins.
- Ras
-
A small G-protein that is part of the Ras superfamily of small G proteins. Members of this family are structurally highly similar and are activated by guanine nucleotide-exchange factors. Ras has an important role in cell-growth control and is mutated in 15% of all human tumours.
- Sos
-
(Son-of-sevenless). Sos is a guanine nucleotide-exchange factor for the Rap-related G-protein Ras.
Rights and permissions
About this article
Cite this article
Rehmann, H., Wittinghofer, A. & Bos, J. Capturing cyclic nucleotides in action: snapshots from crystallographic studies. Nat Rev Mol Cell Biol 8, 63–73 (2007). https://doi.org/10.1038/nrm2082
Issue Date:
DOI: https://doi.org/10.1038/nrm2082
This article is cited by
-
8-Br-cGMP suppresses tumor progression through EGFR/PLC γ1 pathway in epithelial ovarian cancer
Molecular Biology Reports (2024)
-
Differential effects of mutations of POPDC proteins on heteromeric interaction and membrane trafficking
Acta Neuropathologica Communications (2023)
-
The Popeye domain containing gene family encoding a family of cAMP-effector proteins with important functions in striated muscle and beyond
Journal of Muscle Research and Cell Motility (2019)
-
Biochemical Characterization of the Engineered Soluble Photoactivated Guanylate Cyclases from Microbes Expands Optogenetic Tools
Applied Biochemistry and Biotechnology (2018)
-
Modulation of cyclic nucleotide-mediated cellular signaling and gene expression using photoactivated adenylyl cyclase as an optogenetic tool
Scientific Reports (2017)