Abstract
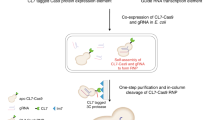
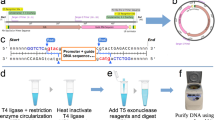
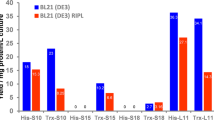
RNA production using in vivo transcription by Escherichia coli allows preparation of milligram quantities of RNA for biochemical, biophysical and structural investigations. We describe here a generic protocol for the overproduction and purification of recombinant RNA using liquid chromatography. The strategy utilizes a transfer RNA (tRNA) as a scaffold that can be removed from the RNA of interest by digestion of the fusion RNA at a designed site by RNase H. The tRNA scaffold serves to enhance the stability and to promote the proper expression of its fusion partners. This protocol describes how to construct a tRNA fusion RNA expression vector; to conduct a pilot experiment to assess the yield of the recombinant RNA both before and after processing of the fusion RNA by RNase H; and to purify the target RNA on a large scale for structural or functional studies. This protocol greatly facilitates production of RNA in a time frame of ∼3 weeks from design to purification. As compared with in vitro methods (transcription, chemical synthesis), this approach is simple, cheap and well suited for large-scale expression and isotope labeling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Marshall, W.S. & Kaiser, R.J. Recent advances in the high-speed solid phase synthesis of RNA. Curr. Opin. Chem. Biol. 8, 222–229 (2004).
Schenborn, E.T. & Mierendorf, R.C. Jr. A novel transcription property of SP6 and T7 RNA polymerases: dependence on template structure. Nucleic Acids Res. 16, 6223–6236 (1985).
Lukavsky, P.J. & Puglisi, J.D. Large-scale preparation and purification of polyacrylamide-free RNA oligonucleotides. RNA 10, 889–893 (2004).
Cheong, H.K., Hwang, E., Lee, C., Choi, B.S. & Cheong, C. Rapid preparation of RNA samples for NMR spectroscopy and X-ray crystallography. Nucleic Acids Res. 32, e84 (2004).
Kieft, J.S. & Batey, R.T. A general method for rapid and nondenaturing purification of RNAs. RNA 10, 988–995 (2004).
Mayer, M. & James, T.L. NMR-based characterisation of phenothiazines as a RNA binding scaffold. J. Am. Chem. Soc. 126, 4453–4460 (2004).
Masson, J.-M & Miller, J.H. Expression of synthetic tRNA genes under the control of a synthetic promoter. Gene 47, 179–183 (1986).
Meinnel, T., Mechulam, Y. & Fayat, G. Fast purification of a functional elongator tRNAmet expressed from a synthetic gene in vivo . Nucleic Acids Res. 16, 8095–8096 (1988).
Tisné, C., Rigourd, M., Marquet, R., Ehresmann, C. & Dardel, F. NMR and biochemical characterization of recombinant human tRNA(Lys)3 expressed in Escherichia coli: identification of posttranscriptional nucleotide modifications required for efficient initiation of HIV-1 reverse transcription. RNA 6, 1403–1412 (2000).
Wallis, N.G., Dardel, F. & Blanquet, S. Heteronuclear NMR studies of the interactions of 15N-labeled methionine-specific transfer RNAs with methionyl-tRNA transformylase. Biochemistry 34, 7668–7677 (1995).
Deutscher, M.P. Ribonucleases, tRNA nucleotidyltransferase, and the 3′ processing of tRNA. Prog. Nucleic Acid Res. Mol. Biol. 39, 209–240 (1990).
Engelke, D.R. & Hopper, A.K. Modified view of tRNA: stability amid sequence diversity. Mol. Cell 21, 144–145 (2006).
Ponchon, L. & Dardel, F. Recombinant RNA technology: the tRNA scaffold. Nat. Methods 4, 571–576 (2007).
Smith, D.B. & Johnson, K.S. Single-step purification of polypeptides expressed in E. coli as fusions with glutathione S-transferase. Gene 67, 31–40 (1988).
McKenna, S.A. et al. Purification and characterization of transcribed RNAs using gel filtration chromatography. Nat. Protoc. 2, 3270–3277 (2007).
Nassal, M. Hepatitis B viruses: reverse transcription a different way. Virus Res. 134, 235–249 (2008).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual 2nd edn. (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, (1989).
Gaudin, C. et al. The tRNA-like domains of E. coli and A. aeolicus transfer-messenger RNA: structural and functional studies. J. Mol. Biol. 331, 457–471 (2003).
Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Fourmy, D., Recht, M.I., Blanchard, S.C. & Puglisi, J.D. Structure of the A site of E. coli 16 ribosomal RNA complexed with an aminoglycoside antibiotic. Science 274, 1367–1371 (1996).
Fürtig, B., Richter, C., Wöhnert, J. & Schwalbe, H. NMR spectroscopy of RNA. Chembiochem. 4, 936–962 (2003).
Schumacher, J., Meyer, N., Riesner, D. & Weidemann, H.L. Diagnostic procedure for detection of viroids and viruses with circular RNAs by 'Return'-gel electrophoresis,. J. Phytopathol. 115, 332–343 (1986).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ponchon, L., Beauvais, G., Nonin-Lecomte, S. et al. A generic protocol for the expression and purification of recombinant RNA in Escherichia coli using a tRNA scaffold. Nat Protoc 4, 947–959 (2009). https://doi.org/10.1038/nprot.2009.67
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.67
This article is cited by
-
A novel recombinant ORF7-siRNA delivered by flexible nano-liposomes inhibits varicella zoster virus infection
Cell & Bioscience (2023)
-
Structures of flavivirus RNA promoters suggest two binding modes with NS5 polymerase
Nature Communications (2021)
-
Branched kissing loops for the construction of diverse RNA homooligomeric nanostructures
Nature Chemistry (2020)
-
Bioengineering of a single long noncoding RNA molecule that carries multiple small RNAs
Applied Microbiology and Biotechnology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.