Abstract
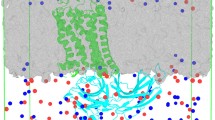
Based on the growing evidence that G-protein coupled receptors (GPCRs) form homo- and hetero-oligomers, models of GPCR signaling are now considering macromolecular assemblies rather than monomers, with the homo-dimer regarded as the minimal oligomeric arrangement required for functional coupling to the G-protein. The dynamic mechanisms of such signaling assemblies are unknown. To gain some insight into properties of GPCR dimers that may be relevant to functional mechanisms, we study their current structural prototype, rhodopsin. We have carried out nanosecond time-scale molecular dynamics (MD) simulations of a rhodopsin dimer and compared the results to the monomer simulated in the same type of bilayer membrane model composed of an equilibrated unit cell of hydrated palmitoyl-oleoyl-phosphatidyl choline (POPC). The dynamic representation of the homo-dimer reveals the location of structural changes in several regions of the monomeric subunits. These changes appear to be more pronounced at the dimerization interface that had been shown to be involved in the activation process [Proc Natl Acad Sci USA 102:17495, 2005]. The results are consistent with a model of GPCR activation that involves allosteric modulation through a single GPCR subunit per dimer.
Similar content being viewed by others
References
Terrillon S, Bouvier M (2004) EMBO Rep 5(1):30
Javitch JA (2004) Mol Pharmacol 66(5):1077
Milligan G (2004) Mol Pharmacol 66(1):1
Bai M (2004) Cell Signal 16(2):175
Park PS, Filipek S, Wells JW, Palczewski K (2004) Biochemistry 43(50):15643
Filizola M, Weinstein H (2005) Curr Opin Drug Discov Devel 8(5):577
Fotiadis D, Jastrzebska B, Philippsen A, Muller DJ, Palczewski K, Engel A (2006) Curr Opin Struct Biol 16(2):252
Guo W, Shi L, Filizola M, Weinstein H, Javitch JA (2005) Proc Natl Acad Sci USA 102(48):17495
Waldhoer M, Fong J, Jones RM, Lunzer MM, Sharma SK, Kostenis E, Portoghese PS, Whistler JL (2005) Proc Natl Acad Sci USA 102(25):9050
Filipek S, Krzysko KA, Fotiadis D, Liang Y, Saperstein DA, Engel A, Palczewski K (2004) Photochem Photobiol Sci 3(6):628
Liang Y, Fotiadis D, Filipek S, Saperstein DA, Palczewski K, Engel A (2003) J Biol Chem 278:21655
Jastrzebska B, Maeda T, Zhu L, Fotiadis D, Filipek S, Engel A, Stenkamp RE, Palczewski K (2004) J Biol Chem 279(52):54663
Fotiadis D, Liang Y, Filipek S, Saperstein DA, Engel A, Palczewski K (2003) Nature 421(6919):127
Teller DC, Okada T, Behnke CA, Palczewski K, Stenkamp RE (2001) Biochemistry 40:7761
Kota P, Reeves PJ, Rajbhandary UL, Khorana HG (2006) Proc Natl Acad Sci USA 103(9):3054
Pin JP, Kniazeff J, Liu J, Binet V, Goudet C, Rondard P, Prezeau L (2005) Febs J 272(12):2947
Duthey B, Caudron S, Perroy J, Bettler B, Fagni L, Pin JP, Prezeau L (2002) J Biol Chem 277(5):3236
Galvez T, Duthey B, Kniazeff J, Blahos J, Rovelli G, Bettler B, Prezeau L, Pin JP (2001) Embo J 20(9):2152
Margeta-Mitrovic M, Jan YN, Jan LY (2001) Proc Natl Acad Sci USA 98:14643
Goudet C, Kniazeff J, Hlavackova V, Malhaire F, Maurel D, Acher F, Blahos J, Prezeau L, Pin JP (2005) J Biol Chem 280(26):24380
Hlavackova V, Goudet C, Kniazeff J, Zikova A, Maurel D, Vol C, Trojanova J, Prezeau L, Pin JP, Blahos J (2005) Embo J 24(3):499
Stenkamp RE, Teller DC, Palczewski K (2005) Arch Pharm (Weinheim) 338(5–6):209
Weinstein H (2006) AAPS J 7(4):871
Okada T, Fujiyoshi Y, Silow M, Navarro J, Landau EM, Shichida Y (2002) Proc Natl Acad Sci USA 99(9):5982
Hassan SA, Mehler EL, Zhang D, Weinstein H (2003) Proteins 51:109
Li XF, Hassan SA, Mehler EL (2005) Proteins Struct Funct Bioinform 60(3):464
Mehler EL, Periole X, Hassan SA, Weinstein HJ (2002) Comput Aided Mol Des 16(11):841
Okada T, Sugihara M, Bondar AN, Elstner M, Entel P, Buss V (2004) J Mol Biol 342(2):571
Periole X, Ceruso MA, Mehler EL (2004) Biochemistry 43(22):6858
Resat H, Mezei M (1994) J Am Chem Soc 116:7451
Resat H, Mezei M (1996) Biophys J 71(3):1179
Marrone TJ, Resat H, Hodge CN, Chang C-H, McCammon JA (1998) Protein Sci 7:573
Berendsen HJC, van der Spoel D, van Drunen R (1995) Comp Phys Comm 91:43
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) J Comput Chem 26(16):1701
Tajkhorshid E, Baudry J, Schulten K, Suhai S (2000) Biophys J 78(2):683
Tajkhorshid E, Paizs B, Suhai S (1997) J Phys Chem B 101:8021
Tajkhorshid E, Suhai S (1999) J Phys Chem B 103:5581
Nina M, Roux B, Smith J (1995) Biophys J 68:25
Tieleman DP, Berendsen HJ (1998) Biophys J 74(6):2786
Tieleman DP, Forrest LR, Sansom MS, Berendsen HJ (1998) Biochemistry 37(50):17554
Sankararamakrishnan R, Weinstein H (2000) Biophys J 79(5):2331
Sankararamakrishnan R, Weinstein H (2004) J Phys Chem B 108:11802
Sankararamakrishnan R, Weinstein H (2002) J Phys Chem B 106:209
Visiers I, Ballesteros JA, Weinstein H (2002) Meth Enzymol 343:329
Fanelli F, De Benedetti PG (2005) Chem Rev 105(9):3297
Berger O, Edholm O, Jahnig F (1997) Biophys J 72:2002
Darden T, York D, Pederson L (1993) J Chem Phys 98:10089
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) J Comput Chem 18:1463
Berendsen HJC, Postma JPM, Gunsteren WF, Hermans J (1981) Intermolecular forces. Riedel, Dordrecht
van Aalten DM, Amadei A, Linssen AB, Eijsink VG, Vriend G, Berendsen HJ (1995) Proteins 22(1):45
Visiers I, Braunheim BB, Weinstein H (2000) Protein Eng 13(9):603
Mezei M, Filizola, M (2006) J Comp Aid Mol Des. 20(2):97
Pitman MC, Grossfield A, Suits F, Feller SE (2005) J Am Chem Soc 127(13):4576
Saam J, Tajkhorshid E, Hayashi S, Schulten K (2002) Biophys J 83(6):3097
Huber T, Botelho AV, Beyer K, Brown MF (2004) Biophys J 86(4):2078
Faraldo-Gomez JD, Forrest LR, Baaden M, Bond PJ, Domene C, Patargias G, Cuthbertson J, Sansom MS (2004) Proteins 57(4):783
Schlegel B, Sippl W, Holtje HD (2005) J Mol Model (Online) 12(1):49
Crozier PS, Stevens MJ, Forrest LR, Woolf TB (2003) J Mol Biol 333(3):493
Fong SL, Tsin AT, Bridges CD, Liou GI (1982) Methods Enzymol 81:133
Petrache HI, Dodd SW, Brown MF (2000) Biophys J 79:3172
Hayward S, Berendsen HJ (1998) Proteins 30(2):144
Farrens DL, Altenbach C, Yang K, Hubbell WL, Khorana HG (1996) Science 274(5288):768
Borhan B, Souto ML, Imai H, Shichida Y, Nakanishi K (2000) Science 288(5474):2209
Ballesteros JA, Weinstein H (1995) Meth Neurosci 25:366
Acknowledgements
We are grateful to Dr. Marc Ceruso for help in setting up the initial dimer simulations, and Dr. Nathalie Basdevant for help with the essential dynamics analysis. This work was supported by NIH grants DA00060, DA012923 (to HW) and DA020032, DA017976 (to MF) from the National Institute on Drug Abuse.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Filizola, M., Wang, S.X. & Weinstein, H. Dynamic models of G-protein coupled receptor dimers: indications of asymmetry in the rhodopsin dimer from molecular dynamics simulations in a POPC bilayer. J Comput Aided Mol Des 20, 405–416 (2006). https://doi.org/10.1007/s10822-006-9053-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-006-9053-3